- Your cart is empty
- Continue Shopping
TSA-Based Multiplex Immunofluorescence in Spatial Biology: A Practical Guide

TSA-Based Multiplex Immunofluorescence in Spatial Biology: A Practical Guide
- Atlantis Bioscience
- Blog
- Reading Time: 7 minutes
Table of Contents
- Why Spatial Context Matters in Multiplex Immunofluorescence
- The Spatial Biology Toolkit: From mIF to Spatial Omics
- Designing a Spatial Biology Experiment
- Choosing the Right Spatial Platform for Your Biological Question
- Deep Dive: Tyramide Signal Amplification
- TyraMax™: Best in Class Brightness and Photostability for TSA Multiplex IHC
- TSA-based multiplexing Practical Challenges and Strategies
- The Future of TSA-Based Multiplex Immunofluorescence in Spatial Biology
- Conclusion
- References
Modern biology has long relied on techniques that analyse cells in isolation. While bulk and single cell sequencing have provided deep molecular insights, they often miss a crucial dimension: spatial context. In spatial biology, preserving tissue architecture is essential for understanding how cells interact within their native microenvironment.
TSA-based multiplex immunofluorescence (mIF) addresses this limitation by enabling high sensitivity detection of multiple protein targets within intact tissue sections. By combining TSA with sequential staining strategies, researchers can map functional protein expression while maintaining spatial relationships between cells.
Why Spatial Context Matters in Multiplex Immunofluorescence
Traditional techniques such as bulk sequencing and even single-cell sequencing typically require tissue dissociation. While these methods reveal valuable information about gene or protein expression, they disrupt the spatial relationships between cells.
However, in many biological processes, location is as important as molecular identity. For example:
- In Oncology: The proximity of immune cells to cancer cells can determine whether an immune response is effective.
- In Neuroscience: Neuronal function depends on precise spatial organisation.
- In Development: Gradients of signalling molecules guide tissue formation.
Without spatial information, we lose sight of cell–cell interactions, functional niches, and the microenvironmental influences that drive disease progression. TSA-based mIF within a spatial biology framework enables high sensitivity protein mapping while preserving these critical spatial relationships.

The Spatial Biology Toolkit: From mIF to Spatial Omics
Multiplex immunofluorescence (mIF) is a cornerstone technique in spatial biology, enabling detection of multiple protein markers on a single tissue section while preserving tissue architecture. In many high-sensitivity workflows, mIF incorporates Tyramide Signal Amplification, often referred to as TSA-based mIF, to enhance signal intensity and enable reliable detection of low-abundance protein targets.
While mIF provides high-resolution protein-level insight, spatial biology encompasses a broader toolkit often referred to as spatial omics. These technologies enable molecular mapping at scale within intact tissues, including:
- Spatial transcriptomics — Capturing the whole transcriptome (10,000 genes) in situ
- Spatial proteomics — Quantifying proteins via antibodies or mass spectrometry
- Multi-omic platforms — Integrating RNA, protein, metabolites, and epigenetic information simultaneously
- Computational spatial mapping — Using AI to reconstruct cellular neighbourhoods and inferring interactions
Together, these approaches provide complementary layers of information. Spatial transcriptomics supports unbiased discovery, whereas TSA-based mIF delivers high-sensitivity protein validation within preserved tissue architecture. This multi-layered framework allows researchers to interpret molecular programmes in the context of cellular organisation and microenvironmental structure.
Designing a Spatial Biology Experiment
Careful experimental design is critical for generating reliable spatial biology data, particularly when implementing TSA-based mIF workflows.
- Define the biological question – Identify whether the focus is immune microenvironments, tumour heterogeneity, tissue development, or disease mechanisms.
- Marker selection – Choose markers that define cell types, functional states, or signalling pathways.
- Tissue preparation – Different tissue types may require FFPE, fresh frozen, or specialised clearing techniques for imaging.
- Technology selection – Decide between mIF, imaging mass cytometry, spatial transcriptomics, or multi-omic platforms depending on required resolution, throughput, and molecular coverage.
- Data analysis planning – Spatial datasets are complex, often requiring advanced computational pipelines for segmentation, clustering, and interaction mapping.

Choosing the Right Spatial Platform for Your Biological Question
Selecting the right spatial biology platform is often the most challenging step in experimental design. It is tempting to pursue the highest plex method available, but the most appropriate technology is the one that answers your specific biological question with sufficient resolution, sensitivity, throughput, and reproducibility.
While this guide focuses on TSA-based multiplex immunofluorescence (mIF), it is essential to understand how it compares to other leading spatial technologies. The table below outlines common scenarios to help determine when TSA-mIF is the optimal choice—and when alternatives may be better suited.
| Technology | Best Used For | Plex Capacity | Sensitivity | Why Choose TSA-mIF Instead? |
| TSA-based mIF | Validating 4–10 markers; high-sensitivity detection of “dim” targets. | 4–10 | Highest (10-100x boost) | Cost-effective. Uses standard lab microscopes; covalent signal allows stripping without data loss. |
| PhenoCycler (CODEX) | High-plex discovery; mapping 40+ cell types in a single tissue. | 40–60+ | Moderate | Less Complexity. PhenoCycler requires specialised fluidics and expensive barcodes; TSA is better for low-abundance proteins. |
| Imaging Mass Cytometry (IMC) | Autofluorescence-heavy tissues (Lung/Liver); absolute quantification. | 40+ | Low to Moderate | Higher Resolution. IMC resolution is often limited to 1µm; TSA-mIF offers crisp subcellular detail and much higher signal. |
| Spatial | Whole-transcriptome discovery; RNA-focused questions. | 1,000–10,000+ | Variable | Protein is King. RNA levels don’t always equal protein levels. TSA validates the “functional” protein expression in situ. |
Key Takeaway
TSA-based mIF is the workhorse for hypothesis-driven protein imaging. It is the only method listed that combines (a) FFPE compatibility, (b) single-molecule sensitivity, (c) standard laboratory equipment, and (d) established utility in high-impact translational research.
If your experiment requires 40+ markers or unbiased discovery, consider CODEX or spatial transcriptomics. If you need to visualise 4–10 carefully selected proteins with high sensitivity and reproducibility, TSA-mIF is the superior choice.
Deep Dive: Tyramide Signal Amplification
As we move toward higher-plex imaging, traditional staining methods often struggle with weak signals. Tyramide Signal Amplification (TSA) has emerged as a key technology to bridge this gap.
Note on scope: Spatial omics encompasses RNA, protein, and metabolite mapping. This section focuses specifically on multiplexed protein imaging, where sensitivity is often the limiting factor. While spatial transcriptomics captures the blueprint of gene expression, spatial proteomics reveals the functional state of cells—post-translational modifications, activated signalling pathways, and therapeutic target engagement. TSA is the critical enabling chemistry for this domain.
How TSA Works
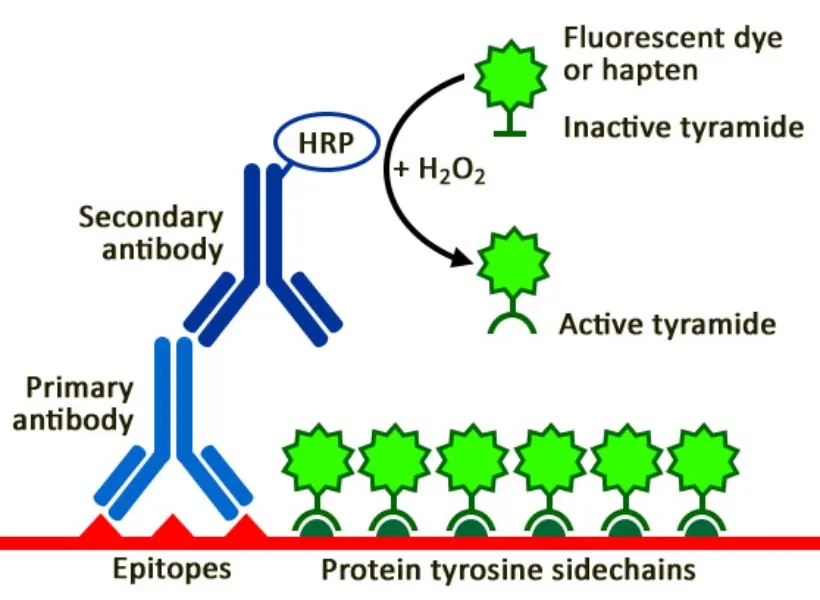
TSA is an enzyme-mediated detection method that uses Horseradish Peroxidase (HRP) to catalyse the deposition of labelled tyramide molecules. These reactive radicals bind covalently to nearby tyrosine residues on the tissue.
The result? A signal that is 10–100 times stronger than conventional IHC. Because the bond is covalent, the fluorescent signal stays “locked” in place even if the primary antibodies are stripped away to make room for the next round of markers.
Why TSA is Essential for Multiplexing
- High Sensitivity: It enables the detection of low-abundance or “dim” markers that would otherwise be lost in background noise.
- Sequential Multiplexing: Researchers can stain, strip the antibodies, and repeat. The TSA signal remains, allowing for the visualisation of dozens of markers on one slide.
- Spatial Precision: The radicals diffuse only a tiny distance before binding, ensuring clear cell boundaries and minimal “bleed” between structures.
TyraMax™: Best in Class Brightness and Photostability for TSA Multiplex IHC
Choosing the right amplification chemistry is the most critical decision in panel development. While several commercial options exist, researchers prioritise brightness, photostability, and ease of automation.
When evaluating the best TSA kits, Biotium offers standout tailored for specific research needs:
TyraMax™ Amplification Dyes and Kits
Compared with commonly used tyramide systems such as Opal and Aluora, TyraMax™ is engineered to deliver higher fluorescence intensity and improved resistance to photobleaching during repeated imaging and stripping cycles. This enhanced performance supports clearer detection of low-abundance protein targets and more stable signal retention in high-plex spatial imaging applications.
Importantly, TyraMax™ reagents remain stable in amplification buffer for up to 24 hours, making them well suited for automated staining platforms where reagent stability and consistency are essential for reproducible multiplex workflows.
By selecting kits that offer high signal-to-noise ratios and chemical stability, labs can significantly reduce the “trial and error” often associated with high-plex TSA workflows.
TSA-based multiplexing Practical Challenges and Strategies
Despite its strengths, TSA-based multiplexing presents practical challenges:
- Complexity and manual labour – Multiple staining, washing, and amplification cycles make the workflow labour-intensive and prone to variation.
- Extended protocol times – Sequential staining cycles increase total assay duration.
- High reagent consumption – Repeated amplification cycles require large amounts of antibodies and tyramide reagents, raising cost and optimisation effort.
- Signal crosstalk or bleed-through – High-plex workflows can experience overlapping fluorescence if not carefully planned.
- Antigen degradation or masking – Repeated retrieval and stripping steps can damage sensitive tissue epitopes.
Strategies to overcome these challenges include:
- Automated and standardised workflows – Reduce manual handling, improve reproducibility, and accelerate staining cycles.
- Protocol optimisation – Careful antibody titration, strategic marker staining order, and fine-tuning tyramide concentration.
- Advanced imaging planning – Selecting fluorophores with minimal spectral overlap and balancing high- versus low-abundance markers
Well-optimised TSA workflows enable reliable, reproducible, and high-resolution spatial maps of protein expression within tissues.
The Future of TSA-Based Multiplex Immunofluorescence in Spatial Biology
The future of spatial biology lies in integration. Advances in AI-driven image analysis are enabling correlation of a cell’s genetic state with its physical location and neighbouring proteomic profile within intact tissue. As TSA-based multiplex immunofluorescence workflows become more automated and higher-throughput, high-sensitivity spatial protein mapping is increasingly positioned to support translational research and clinical diagnostics.

Conclusion
Spatial biology is transforming our understanding of life by preserving the “neighbourhood” in which cells live and work. By leveraging the sensitivity of TSA and the power of multiplex imaging, researchers are finally seeing the full picture—accelerating discoveries that will define the next generation of precision medicine.
References
Bressan D, Battistoni G, Hannon GJ. The dawn of spatial omics. Science. 2023 Aug 4;381(6657):eabq4964. doi: 10.1126/science.abq4964.
Liu Y, Dai Y, Wang L. Spatial omics at the forefront: emerging technologies, analytical innovations, and clinical applications. Cancer Cell. 2026 Jan 12;44(1):24-49. doi: 10.1016/j.ccell.2025.12.009.
Moffitt JR, Lundberg E, Heyn H. The emerging landscape of spatial profiling technologies. Nat Rev Genet. 2022 Dec;23(12):741-759. doi: 10.1038/s41576-022-00515-3.
Radosevic-Robin N, Kossai M, Penault-Llorca F. New-generation technologies for spatial tissue analysis, indispensable tools for deciphering intratumor heterogeneity in the development of antibody-drug conjugates and radio-immunoconjugates for cancer treatment. Transl Breast Cancer Res. 2023 Sep 20;4:28. doi: 10.21037/tbcr-23-38.
Sheng W, Zhang C, Mohiuddin TM, Al-Rawe M, Zeppernick F, Falcone FH, Meinhold-Heerlein I, Hussain AF. Multiplex Immunofluorescence: A Powerful Tool in Cancer Immunotherapy. Int J Mol Sci. 2023 Feb 4;24(4):3086. doi: 10.3390/ijms24043086.
Stack EC, Wang C, Roman KA, Hoyt CC. Multiplexed immunohistochemistry, imaging, and quantitation: a review, with an assessment of Tyramide signal amplification, multispectral imaging and multiplex analysis. Methods. 2014 Nov;70(1):46-58. doi: 10.1016/j.ymeth.2014.08.016.
Tan WCC, Nerurkar SN, Cai HY, Ng HHM, Wu D, Wee YTF, Lim JCT, Yeong J, Lim TKH. Overview of multiplex immunohistochemistry/immunofluorescence techniques in the era of cancer immunotherapy. Cancer Commun (Lond). 2020 Apr;40(4):135-153. doi: 10.1002/cac2.12023.
CONTACT

QUESTIONS IN YOUR MIND?
Connect With Our Technical Specialist.

KNOW WHAT YOU WANT?
Request For A Quotation.
OTHER BLOGS YOU MIGHT LIKE
HOW CAN WE HELP YOU? Our specialists are to help you find the best product for your application. We will be happy to help you find the right product for the job.

TALK TO A SPECIALIST
Contact our Customer Care, Sales & Scientific Assistance

EMAIL US
Consult and asked questions about our products & services

DOCUMENTATION
Documentation of Technical & Safety Data Sheet, Guides and more...

