Summary
HRM analysis stands out as a dependable and streamlined approach for genetic variation analysis. Its adaptability, sensitivity, and cost-effectiveness render it an invaluable asset in identifying and characterising genetic variations across diverse domains, spanning medical genetics, cancer research, microbiology, and plant genetics. As technological advancements persist and applications broaden, HRM analysis is poised to remain a pivotal player in the molecular diagnostics arena. Its ongoing contributions are anticipated to drive advancements in personalised medicine, ultimately leading to enhanced patient outcomes.
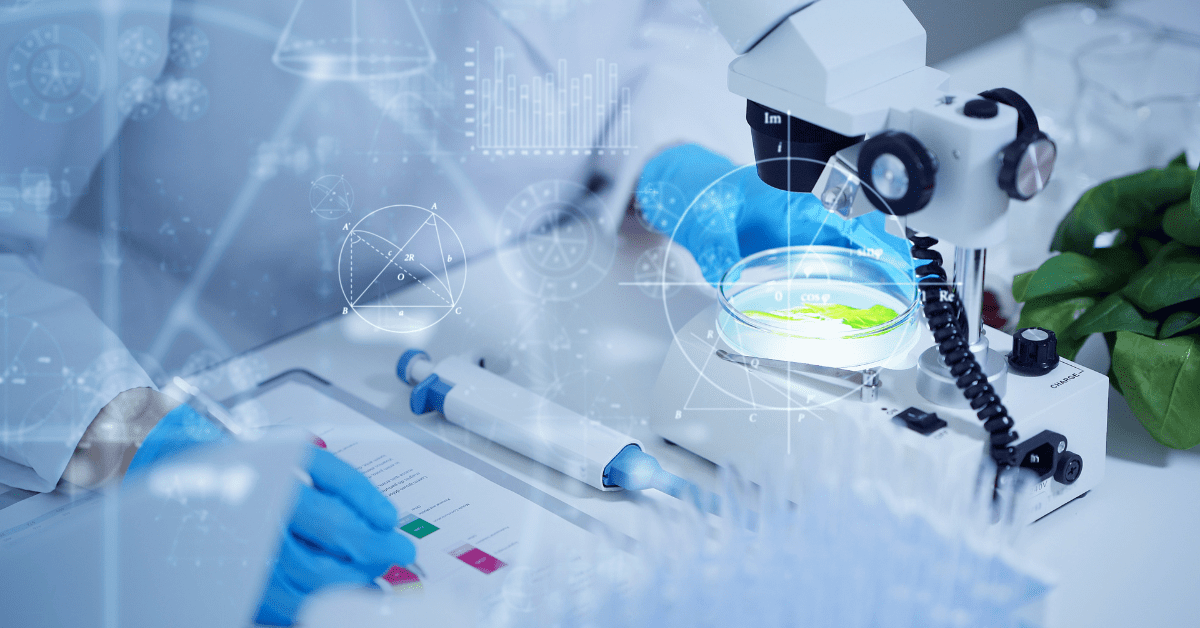
High-Resolution Melting (HRM) analysis is a post-PCR technique used in molecular biology to distinguish variations in DNA sequences. It is a simple and fast method that is based on the melting behaviour of the PCR products. It is particularly useful for detecting genetic variations, such as single nucleotide polymorphisms (SNPs) or mutations, in DNA samples. HRM analysis is often employed in various applications, including genotyping, mutation scanning, and methylation analysis.
What Makes HRM Analysis Particularly Advantageous?
HRM analysis offers significant benefits compared to alternative genotyping technologies like sequencing and Taqman SNP typing. Notably, it is a more cost-effective solution, making it particularly suitable for large-scale genotyping projects. With high sensitivity, it can detect even a single nucleotide change. The workflow is rapid, enabling accurate genotyping of extensive sample sets within a short timeframe. This straightforward assay eliminates the need for specific probes or post-PCR processing steps and can be conducted in any laboratory equipped with an HRM-capable real-time PCR machine.
Here’s a Brief Overview of How HRM Analysis Works:
The process typically begins with PCR amplification of the target DNA region in which their mutation of interest lies in the presence of a double-stranded (dsDNA)-binding dye. This binding dye exhibits high fluorescence when attached to dsDNA and low fluorescence when unbound. Following amplification, the resulting amplicon undergoes a gradual temperature increase, leading to the separation (melting) of dsDNA strands. With rising temperature, the DNA denatures, and the dsDNA transforms into single strands. As this separation occurs, the fluorescence emitted by the dye bound to the dsDNA progressively diminishes.
The outcome is a distinctive melt curve profile characteristic of the amplicon. The term “high resolution” in HRM analysis pertains to the precise monitoring of fluorescence changes during the temperature increase. The DNA melting profile is captured with heightened sensitivity, enabling the detection of subtle distinctions in the melting behaviour of DNA sequences. The recorded melting curve is then analysed to discern variations in the DNA sequence.
Mutations or polymorphisms can influence the melting characteristics of DNA. A comparative analysis of the sample’s melting curve with those of known reference samples can unveil the presence of genetic variations.
Elevating HRM Analysis Excellence with EvaGreen® Dye
For successful HRM analysis, a saturating concentration of dsDNA-binding dyes is essential, striking a balance that does not inhibit the PCR. The quantity of fluorescent dye bound to dsDNA can vary based on factors such as amplicon length, composition, and PCR conditions. Ideally, determining the optimal dye concentration involves empirical testing through a titration experiment.
EvaGreen® dye is a stellar choice for HRM analysis. This green, fluorescent nucleic acid dye is spectrally similar to fluorescein (FAM) or SYBR® dye Green I when bound to dsDNA, making the dye readily compatible with instruments equipped with the 488 nm argon laser or any visible light excitation with wavelength in the region. EvaGreen® dye stands out for its exceptional thermal and hydrolytic stability, offering convenience in routine handling.
While inherently nonfluorescent, EvaGreen® dye becomes highly fluorescent upon binding to dsDNA. Its non-mutagenic and non-cytotoxic nature and impermeability to cell membranes, sets it apart from SYBR Green I, a mutation enhancer that enter cells rapidly.
The distinctive properties of EvaGreen® dye have positioned it as a key player in quantitative real-time PCR (qPCR) applications and HRM analysis. In comparison to the widely used SYBR Green I, EvaGreen® dye tends to be less inhibitory to PCR and less prone to causing nonspecific amplification. This advantage allows EvaGreen® dye to be employed at a higher concentration, leading to a more robust PCR signal and high-resolution DNA melt analysis.
*Note: HRM is a registered trademark of Idaho Technologies, Inc./BioFire Defense, LLC
What are the Applications for HRM Analysis:
HRM analysis finds applications across various fields of molecular biology and genetics. Here’s a list of some common applications:
Genotyping:
HRM analysis is widely used for genotyping, which involves identifying genetic variations, such SNPs, insertions, deletions, or strand complementarity (such as heterozygous or homozygous material), within DNA samples. It enables researchers to distinguish between different alleles or genotypes based on their melting profiles.
Mutation Scanning:
HRM analysis facilitates mutation scanning, a process used to screen large DNA regions or gene panels for the presence of mutations or sequence variations. It allows researchers to efficiently identify potential mutations that may be associated with diseases or genetic disorders. It is often used in profiling genetic alterations in tumour samples, for the detection of mutations associated with cancer.
Methylation Analysis:
DNA methylation, the addition of methyl groups to cytosine residues, plays a critical role in gene regulation and epigenetic modifications. Methylated DNA and unmethylated DNA acquire different sequences after bisulphite treatment resulting in PCR products with markedly different melting profiles. HRM analysis can be used to study DNA methylation patterns by assessing the melting behaviour of methylated and unmethylated DNA sequences. The PCR reaction products can also be sequenced to elucidate specific positions of DNA methylation.
In a recent publication in Molecular Cancer, Kim et al. introduced the methylation-sensitive high-resolution analysis (MS-HRM) assay for the diagnosis of hepatocellular cancer (HCC) in cell-free DNA (cfDNA). Through the application of this MS-HRM method, they successfully identified HCC in liquid biopsy samples, surpassing the performance of the alpha-fetoprotein (AFP) test. These results suggest the potential use of a straightforward PCR-based technique for the diagnostic detection of HCC.
Pathogen Detection:
HRM analysis is employed in the detection and identification of pathogens, including viruses, bacteria, and fungi. By targeting specific genomic regions or conserved sequences, HRM can help distinguish between different pathogen strains or detect the presence of infectious agents in clinical or environmental samples.
In a recent publication in Analytical Chemistry, Pei-Wei Lee et al. introduced an innovative method that seamlessly combines digital PCR and HRM® techniques, surpassing the capabilities of conventional approaches utilising EvaGreen® Dye for enhanced single-cell resolution. The authors successfully implemented this novel workflow, resulting in a digital PCR-HRM assay with significantly improved accuracy in identifying bacteria. This breakthrough not only enhances our understanding of bacterial identification but also sets the stage for the development of future digital PCR-HRM assays targeting a broad spectrum of infectious diseases.
Plant Genetics and Breeding:
In plant genetics and breeding programs, HRM analysis is used for marker-assisted selection, genetic mapping, and diversity analysis. It helps researchers identify and characterise genetic variants associated with desirable traits, such as disease resistance, yield, or quality attributes.
References:
Erali M, Voelkerding KV, Wittwer CT. High resolution melting applications for clinical laboratory medicine. Exp Mol Pathol. 2008;85(1):50-58. doi:10.1016/j.yexmp.2008.03.012
Garritano S, Gemignani F, Voegele C, et al. Determining the effectiveness of High Resolution Melting analysis for SNP genotyping and mutation scanning at the TP53 locus. BMC Genet. 2009;10:5. Published 2009 Feb 17. doi:10.1186/1471-2156-10-5
Kim SC, Kim DW, Cho EJ, et al. A circulating cell-free DNA methylation signature for the detection of hepatocellular carcinoma. Mol Cancer. 2023;22(1):164. Published 2023 Oct 6. doi:10.1186/s12943-023-01872-1
Lee PW, Chen L, Hsieh K, Traylor A, Wang TH. Harnessing Variabilities in Digital Melt Curves for Accurate Identification of Bacteria. Anal Chem. 2023;95(42):15522-15530. doi:10.1021/acs.analchem.3c01654
Ohta T, Tokishita S, Yamagata H. Ethidium bromide and SYBR Green I enhance the genotoxicity of UV- doi:10.1016/s1383-5718(01)00155-3irradiation and chemical mutagens in E. coli. Mutat Res. 2001;492(1-2):91-97. doi:10.1016/s1383-5718(01)00155-3
Reed GH, Kent JO, Wittwer CT. High-resolution DNA melting analysis for simple and efficient molecular diagnostics. Pharmacogenomics. 2007;8(6):597-608. doi:10.2217/14622416.8.6.597
Vossen RH, Aten E, Roos A, den Dunnen JT. High-resolution melting analysis (HRMA): more than just sequence variant screening. Hum Mutat. 2009;30(6):860-866. doi:10.1002/humu.21019 Wojdacz TK, Dobrovic A. Methylation-sensitive high resolution melting (MS-HRM): a new approach for sensitive and high-throughput assessment of methylation. Nucleic Acids Res. 2007;35(6):e41. doi:10.1093/nar/gkm013